4f Edge Detection — Learning the Vortex Phase Filter#
This notebook is the 4f extension notebook. It reuses the same 4f geometry from 4f_correlator.ipynb, but changes the task: instead of using a fixed matched filter for template matching, we learn a phase-only Fourier-plane filter for edge detection.
The optimizer is trained end-to-end on synthetic binary scenes, with no hard-coded analytic solution. The goal is for it to independently recover the spiral (vortex) phase filter \(\phi(x, y) = \arg(x + iy)\), the standard phase-only Fourier filter for isotropic edge enhancement.
Assumes you know#
the 4f optical path and sampling-matched focal length from 4f_correlator.ipynb,
why the Fourier plane is the natural place to apply a spatial-frequency filter, and
basic gradient-based optimization in JAX.
New ideas in this notebook#
generating a compact training set of random binary scenes and edge targets,
parameterizing a phase-only Fourier-plane mask with a bounded phase map,
training the 4f system end-to-end on an image-processing objective, and
comparing the learned phase against the analytical vortex solution.
Relation to the 4f correlator notebook#
The optical layout is the same as in 4f_correlator.ipynb. This notebook does not repeat the full derivation of the 4f geometry; it focuses on what changes when the Fourier-plane element becomes trainable.
0 Imports#
We use JAX and Optax for differentiable optimization, together with the same fouriax
optical components used in the correlator notebook.
from __future__ import annotations
from pathlib import Path
import jax
import jax.numpy as jnp
import matplotlib.pyplot as plt
import numpy as np
import optax
import fouriax as fx
%matplotlib inline
EXAMPLES_ROOT = Path.cwd() / "examples"
EXAMPLES_ARTIFACTS_DIR = EXAMPLES_ROOT / "artifacts"
1 Paths and Parameters#
Artifact outputs are written under artifacts/, including the sampled training-scene
preview figure, optimization summary JSON, and final comparison plot.
The main parameters now control both the optical system and the learning problem: grid and wavelength settings define the 4f geometry, while epoch count, learning rate, and dataset sizes define the optimization workload.
ARTIFACTS_DIR = Path(str(EXAMPLES_ARTIFACTS_DIR))
PLOT_PATH = ARTIFACTS_DIR / "4f_edge_optimization.png"
SUMMARY_PATH = ARTIFACTS_DIR / "4f_edge_optimization_summary.json"
SEED = 0
WAVELENGTH_UM = 0.532
N_MEDIUM = 1.0
GRID_N = 128
GRID_DX_UM = 2.0
EPOCHS = 25
LR = 0.005
N_TRAIN_SCENES = 1000
N_TEST_SCENES = 100
BATCH_SIZE = 8
PLOT = True
2 Helper Functions#
The helper functions define the synthetic data model used for training and the analytical vortex phase used later for comparison. The 4f derivation itself is assumed from 4f_correlator.ipynb.
def random_scene(key: jax.Array, grid: fx.Grid) -> jnp.ndarray:
noise = jax.random.normal(key, grid.shape)
k = jnp.fft.fftn(noise, axes=(-2, -1))
freq_x, freq_y = grid.frequency_grid()
sigma_freq = 1.0 / (32.0 * grid.dx_um)
lpf = jnp.exp(-(freq_x**2 + freq_y**2) / (2 * sigma_freq**2))
smooth = jnp.real(jnp.fft.ifftn(k * lpf, axes=(-2, -1)))
return (smooth > 0).astype(jnp.float32)
def edge_target(scene: jnp.ndarray) -> jnp.ndarray:
padded = jnp.pad(scene, 1, mode="edge")
gx = padded[1:-1, 2:] - padded[1:-1, :-2]
gy = padded[2:, 1:-1] - padded[:-2, 1:-1]
mag = jnp.sqrt(gx**2 + gy**2)
return mag / jnp.maximum(jnp.max(mag), 1e-12)
def sampling_matched_f(grid: fx.Grid) -> float:
return N_MEDIUM * grid.nx * grid.dx_um**2 / WAVELENGTH_UM
def analytical_spiral_phase(grid: fx.Grid) -> jnp.ndarray:
x, y = grid.spatial_grid()
return jnp.arctan2(y, x) + jnp.pi # [0, 2π], centered on optical axis
def make_test_scene(grid: fx.Grid) -> jnp.ndarray:
x, y = grid.spatial_grid()
half = grid.nx * grid.dx_um / 2.0
scene = jnp.zeros(grid.shape, dtype=jnp.float32)
scene = scene + (
(jnp.abs(x - 0.2 * half) < 0.15 * half) & (jnp.abs(y + 0.1 * half) < 0.15 * half)
).astype(jnp.float32)
scene = scene + (
(jnp.abs(x + 0.3 * half) < 0.1 * half) & (jnp.abs(y - 0.25 * half) < 0.1 * half)
).astype(jnp.float32)
r = jnp.sqrt((x + 0.1 * half) ** 2 + (y + 0.3 * half) ** 2)
scene = scene + (r < 0.12 * half).astype(jnp.float32)
return jnp.clip(scene, 0.0, 1.0)
def measure_scenes(
module: fx.OpticalModule,
scenes: jnp.ndarray,
grid: fx.Grid,
spectrum: fx.Spectrum,
) -> jnp.ndarray:
scenes = jnp.asarray(scenes, dtype=jnp.float32)
if scenes.ndim == 2:
scenes = scenes[None, :, :]
field_in = fx.Field(
data=scenes[:, None, :, :].astype(jnp.complex64),
grid=grid,
spectrum=spectrum,
)
return module.measure(field_in)[..., ::-1, ::-1]
3 Setup#
The optical layout is identical to the correlator notebook except for the Fourier-plane element: instead of a fixed complex matched filter, we insert a trainable phase-only mask.
The system follows the same geometry as the correlator notebook:
The phase \(\phi\) is the only trainable parameter. We parameterise it as \(\phi = 2\pi\,\sigma(\theta)\) where \(\sigma\) is the sigmoid function and \(\theta\) is an unconstrained array optimised by Adam.
This bounded parameterization preserves the phase-only constraint while keeping the optimizer in an unconstrained parameter space.
grid = fx.Grid.from_extent(nx=GRID_N, ny=GRID_N, dx_um=GRID_DX_UM, dy_um=GRID_DX_UM)
spectrum = fx.Spectrum.from_scalar(WAVELENGTH_UM)
f_um = sampling_matched_f(grid)
prop = fx.ASMPropagator(
distance_um=f_um,
use_sampling_planner=False,
warn_on_regime_mismatch=False,
)
lens = fx.ThinLens(focal_length_um=f_um)
def build_module(raw_phase: jnp.ndarray) -> fx.OpticalModule:
phase = 2.0 * jnp.pi * jax.nn.sigmoid(raw_phase)
return fx.OpticalModule(
layers=(
prop,
lens,
prop,
fx.ComplexMask(phase_map_rad=phase),
prop,
lens,
prop,
),
sensor=fx.DetectorArray(detector_grid=grid),
)
4 Training Data#
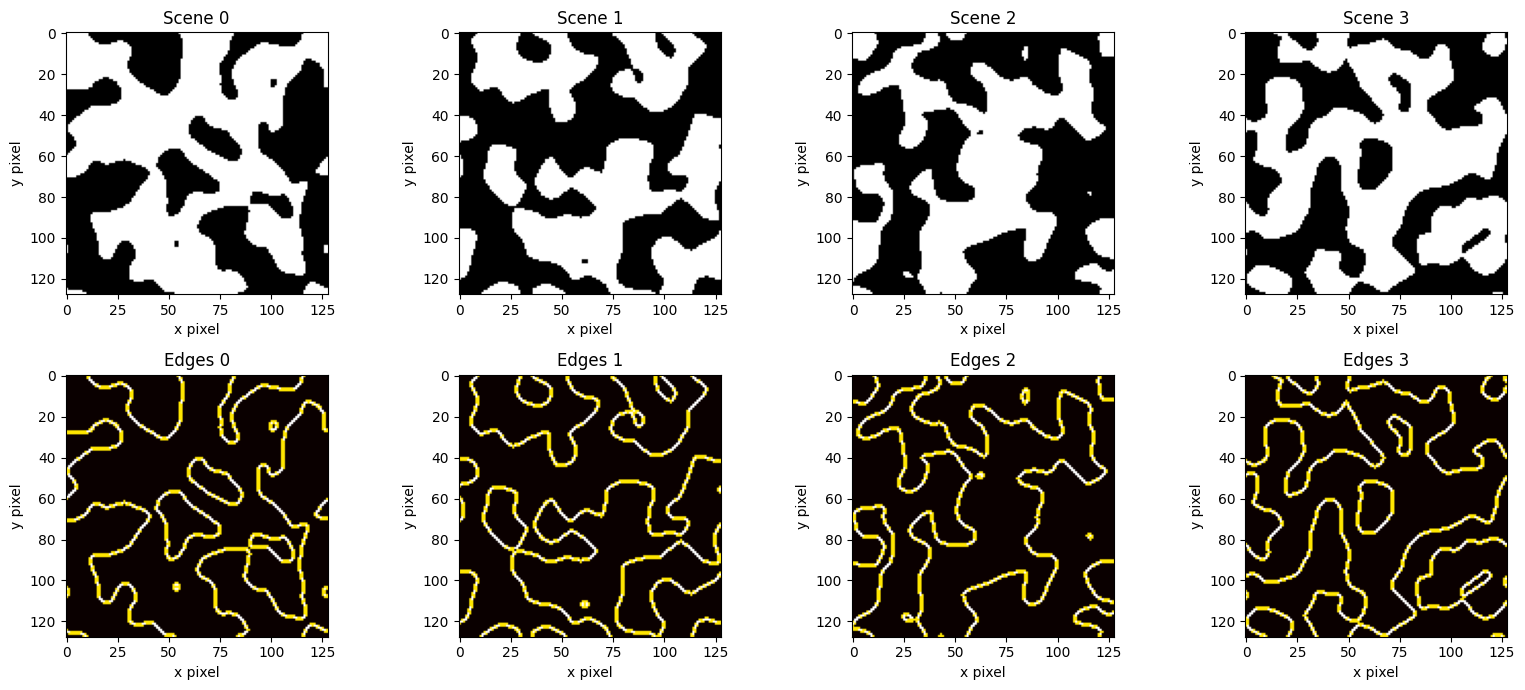
Each training example is simple enough to generate procedurally, while still containing edges at many orientations and locations.
Each training scene is generated by:
Drawing white noise on the simulation grid.
Low-pass filtering in \(k\)-space to produce smooth blobs.
Thresholding at zero to obtain a binary scene.
The smoothing scale controls feature size; larger values yield bigger blobs with fewer, longer edges. The corresponding edge targets are computed via central finite differences (a discrete gradient magnitude).
This keeps the learning problem focused on a single question: can a trainable 4f Fourier-plane phase filter learn a rotationally symmetric edge detector from data alone?
key = jax.random.PRNGKey(SEED)
key, *train_keys = jax.random.split(key, N_TRAIN_SCENES + 1)
train_scenes = jnp.stack([random_scene(k, grid) for k in train_keys])
train_targets = jnp.stack([edge_target(s) for s in train_scenes])
key, *test_keys = jax.random.split(key, N_TEST_SCENES + 1)
test_scenes = jnp.stack([random_scene(k, grid) for k in test_keys])
test_targets = jnp.stack([edge_target(s) for s in test_scenes])
if PLOT:
fig, axes = plt.subplots(2, 4, figsize=(16, 7))
for col in range(4):
axes[0, col].imshow(np.asarray(train_scenes[col]), cmap="gray")
axes[0, col].set_title(f"Scene {col}")
axes[1, col].imshow(np.asarray(train_targets[col]), cmap="hot")
axes[1, col].set_title(f"Edges {col}")
for ax in axes.flat:
ax.set_xlabel("x pixel")
ax.set_ylabel("y pixel")
fig.tight_layout()
save_path = ARTIFACTS_DIR / "4f_edge_optimization_scenes.png"
fig.savefig(save_path)
plt.show()
W0407 20:57:51.292117 1079429 cpp_gen_intrinsics.cc:74] Empty bitcode string provided for eigen. Optimizations relying on this IR will be disabled.
5 Loss Function and Optimization#
The loss is the mean squared error between the normalized 4f output intensity and the target edge map. Normalization removes overall scale as a shortcut, so the optimization has to match spatial structure rather than just total brightness.
Training proceeds over a procedurally generated dataset of scenes and targets. We monitor validation loss on a held-out subset to check whether the learned filter generalizes beyond the individual scenes seen during optimization.
key, init_key = jax.random.split(key)
raw_phase = 0.1 * jax.random.normal(init_key, (grid.ny, grid.nx))
optimizer = optax.adam(LR)
n_test_eval = min(10, N_TEST_SCENES) # evaluate on a subset for speed
val_data = (test_scenes[:n_test_eval], test_targets[:n_test_eval])
def batch_loss_fn(
params: jnp.ndarray,
batch: tuple[jnp.ndarray, jnp.ndarray],
) -> jnp.ndarray:
scenes, targets = batch
module = build_module(params)
out = measure_scenes(module, scenes, grid, spectrum)
out_n = out / jnp.maximum(jnp.max(out, axis=(-2, -1), keepdims=True), 1e-12)
return jnp.mean((out_n - jnp.asarray(targets, dtype=jnp.float32)) ** 2)
result = fx.optim.optimize_dataset_optical_module(
init_params=raw_phase,
build_module=build_module,
batch_loss_fn=batch_loss_fn,
optimizer=optimizer,
train_data=(train_scenes, train_targets),
batch_size=BATCH_SIZE,
epochs=EPOCHS,
val_data=val_data,
seed=SEED,
)
train_history = result.params_result.train_loss_history
test_history = [
(record.step, record.metrics["val_loss"]) for record in result.params_result.val_history
]
A module that was compiled using NumPy 1.x cannot be run in
NumPy 2.4.1 as it may crash. To support both 1.x and 2.x
versions of NumPy, modules must be compiled with NumPy 2.0.
Some module may need to rebuild instead e.g. with 'pybind11>=2.12'.
If you are a user of the module, the easiest solution will be to
downgrade to 'numpy<2' or try to upgrade the affected module.
We expect that some modules will need time to support NumPy 2.
Traceback (most recent call last): File "<frozen runpy>", line 198, in _run_module_as_main
File "<frozen runpy>", line 88, in _run_code
File "/opt/anaconda3/lib/python3.12/site-packages/ipykernel_launcher.py", line 17, in <module>
app.launch_new_instance()
File "/opt/anaconda3/lib/python3.12/site-packages/traitlets/config/application.py", line 1075, in launch_instance
app.start()
File "/opt/anaconda3/lib/python3.12/site-packages/ipykernel/kernelapp.py", line 701, in start
self.io_loop.start()
File "/opt/anaconda3/lib/python3.12/site-packages/tornado/platform/asyncio.py", line 205, in start
self.asyncio_loop.run_forever()
File "/opt/anaconda3/lib/python3.12/asyncio/base_events.py", line 641, in run_forever
self._run_once()
File "/opt/anaconda3/lib/python3.12/asyncio/base_events.py", line 1986, in _run_once
handle._run()
File "/opt/anaconda3/lib/python3.12/asyncio/events.py", line 88, in _run
self._context.run(self._callback, *self._args)
File "/opt/anaconda3/lib/python3.12/site-packages/ipykernel/kernelbase.py", line 534, in dispatch_queue
await self.process_one()
File "/opt/anaconda3/lib/python3.12/site-packages/ipykernel/kernelbase.py", line 523, in process_one
await dispatch(*args)
File "/opt/anaconda3/lib/python3.12/site-packages/ipykernel/kernelbase.py", line 429, in dispatch_shell
await result
File "/opt/anaconda3/lib/python3.12/site-packages/ipykernel/kernelbase.py", line 767, in execute_request
reply_content = await reply_content
File "/opt/anaconda3/lib/python3.12/site-packages/ipykernel/ipkernel.py", line 429, in do_execute
res = shell.run_cell(
File "/opt/anaconda3/lib/python3.12/site-packages/ipykernel/zmqshell.py", line 549, in run_cell
return super().run_cell(*args, **kwargs)
File "/opt/anaconda3/lib/python3.12/site-packages/IPython/core/interactiveshell.py", line 3075, in run_cell
result = self._run_cell(
File "/opt/anaconda3/lib/python3.12/site-packages/IPython/core/interactiveshell.py", line 3130, in _run_cell
result = runner(coro)
File "/opt/anaconda3/lib/python3.12/site-packages/IPython/core/async_helpers.py", line 128, in _pseudo_sync_runner
coro.send(None)
File "/opt/anaconda3/lib/python3.12/site-packages/IPython/core/interactiveshell.py", line 3334, in run_cell_async
has_raised = await self.run_ast_nodes(code_ast.body, cell_name,
File "/opt/anaconda3/lib/python3.12/site-packages/IPython/core/interactiveshell.py", line 3517, in run_ast_nodes
if await self.run_code(code, result, async_=asy):
File "/opt/anaconda3/lib/python3.12/site-packages/IPython/core/interactiveshell.py", line 3577, in run_code
exec(code_obj, self.user_global_ns, self.user_ns)
File "/var/folders/h9/2zpp40gn24l3ddb3fnkq7fzh0000gn/T/ipykernel_50301/2443383685.py", line 18, in <module>
result = fx.optim.optimize_dataset_optical_module(
File "/Users/liam/metasurface/fouriax/src/fouriax/optim.py", line 752, in optimize_dataset_optical_module
params_result = optimize_dataset_params(
File "/Users/liam/metasurface/fouriax/src/fouriax/optim.py", line 584, in optimize_dataset_params
epoch_batches = iter_batches(epoch)
File "/Users/liam/metasurface/fouriax/src/fouriax/optim.py", line 530, in iter_batches
return batch_iter_any(train_data, epoch)
File "/Users/liam/metasurface/fouriax/src/fouriax/optim.py", line 727, in batch_iter_fn
return iter_minibatches(
File "/Users/liam/metasurface/fouriax/src/fouriax/optim.py", line 200, in iter_minibatches
from sklearn.utils import shuffle as sklearn_shuffle
File "/opt/anaconda3/lib/python3.12/site-packages/sklearn/__init__.py", line 84, in <module>
from .base import clone
File "/opt/anaconda3/lib/python3.12/site-packages/sklearn/base.py", line 19, in <module>
from .utils._estimator_html_repr import _HTMLDocumentationLinkMixin, estimator_html_repr
File "/opt/anaconda3/lib/python3.12/site-packages/sklearn/utils/__init__.py", line 11, in <module>
from ._chunking import gen_batches, gen_even_slices
File "/opt/anaconda3/lib/python3.12/site-packages/sklearn/utils/_chunking.py", line 8, in <module>
from ._param_validation import Interval, validate_params
File "/opt/anaconda3/lib/python3.12/site-packages/sklearn/utils/_param_validation.py", line 14, in <module>
from .validation import _is_arraylike_not_scalar
File "/opt/anaconda3/lib/python3.12/site-packages/sklearn/utils/validation.py", line 26, in <module>
from ..utils._array_api import _asarray_with_order, _is_numpy_namespace, get_namespace
File "/opt/anaconda3/lib/python3.12/site-packages/sklearn/utils/_array_api.py", line 11, in <module>
from .fixes import parse_version
File "/opt/anaconda3/lib/python3.12/site-packages/sklearn/utils/fixes.py", line 24, in <module>
import pandas as pd
File "/opt/anaconda3/lib/python3.12/site-packages/pandas/__init__.py", line 26, in <module>
from pandas.compat import (
File "/opt/anaconda3/lib/python3.12/site-packages/pandas/compat/__init__.py", line 27, in <module>
from pandas.compat.pyarrow import (
File "/opt/anaconda3/lib/python3.12/site-packages/pandas/compat/pyarrow.py", line 8, in <module>
import pyarrow as pa
File "/opt/anaconda3/lib/python3.12/site-packages/pyarrow/__init__.py", line 65, in <module>
import pyarrow.lib as _lib
---------------------------------------------------------------------------
TypeError Traceback (most recent call last)
File ~/metasurface/fouriax/src/fouriax/optim.py:527, in optimize_dataset_params.<locals>.iter_batches(epoch)
526 try:
--> 527 return batch_iter_any(train_data, epoch, rng)
528 except TypeError:
TypeError: optimize_dataset_optical_module.<locals>.batch_iter_fn() takes 2 positional arguments but 3 were given
During handling of the above exception, another exception occurred:
ImportError Traceback (most recent call last)
File /opt/anaconda3/lib/python3.12/site-packages/numpy/core/_multiarray_umath.py:46, in __getattr__(attr_name)
41 # Also print the message (with traceback). This is because old versions
42 # of NumPy unfortunately set up the import to replace (and hide) the
43 # error. The traceback shouldn't be needed, but e.g. pytest plugins
44 # seem to swallow it and we should be failing anyway...
45 sys.stderr.write(msg + tb_msg)
---> 46 raise ImportError(msg)
48 ret = getattr(_multiarray_umath, attr_name, None)
49 if ret is None:
ImportError:
A module that was compiled using NumPy 1.x cannot be run in
NumPy 2.4.1 as it may crash. To support both 1.x and 2.x
versions of NumPy, modules must be compiled with NumPy 2.0.
Some module may need to rebuild instead e.g. with 'pybind11>=2.12'.
If you are a user of the module, the easiest solution will be to
downgrade to 'numpy<2' or try to upgrade the affected module.
We expect that some modules will need time to support NumPy 2.
A module that was compiled using NumPy 1.x cannot be run in
NumPy 2.4.1 as it may crash. To support both 1.x and 2.x
versions of NumPy, modules must be compiled with NumPy 2.0.
Some module may need to rebuild instead e.g. with 'pybind11>=2.12'.
If you are a user of the module, the easiest solution will be to
downgrade to 'numpy<2' or try to upgrade the affected module.
We expect that some modules will need time to support NumPy 2.
Traceback (most recent call last): File "<frozen runpy>", line 198, in _run_module_as_main
File "<frozen runpy>", line 88, in _run_code
File "/opt/anaconda3/lib/python3.12/site-packages/ipykernel_launcher.py", line 17, in <module>
app.launch_new_instance()
File "/opt/anaconda3/lib/python3.12/site-packages/traitlets/config/application.py", line 1075, in launch_instance
app.start()
File "/opt/anaconda3/lib/python3.12/site-packages/ipykernel/kernelapp.py", line 701, in start
self.io_loop.start()
File "/opt/anaconda3/lib/python3.12/site-packages/tornado/platform/asyncio.py", line 205, in start
self.asyncio_loop.run_forever()
File "/opt/anaconda3/lib/python3.12/asyncio/base_events.py", line 641, in run_forever
self._run_once()
File "/opt/anaconda3/lib/python3.12/asyncio/base_events.py", line 1986, in _run_once
handle._run()
File "/opt/anaconda3/lib/python3.12/asyncio/events.py", line 88, in _run
self._context.run(self._callback, *self._args)
File "/opt/anaconda3/lib/python3.12/site-packages/ipykernel/kernelbase.py", line 534, in dispatch_queue
await self.process_one()
File "/opt/anaconda3/lib/python3.12/site-packages/ipykernel/kernelbase.py", line 523, in process_one
await dispatch(*args)
File "/opt/anaconda3/lib/python3.12/site-packages/ipykernel/kernelbase.py", line 429, in dispatch_shell
await result
File "/opt/anaconda3/lib/python3.12/site-packages/ipykernel/kernelbase.py", line 767, in execute_request
reply_content = await reply_content
File "/opt/anaconda3/lib/python3.12/site-packages/ipykernel/ipkernel.py", line 429, in do_execute
res = shell.run_cell(
File "/opt/anaconda3/lib/python3.12/site-packages/ipykernel/zmqshell.py", line 549, in run_cell
return super().run_cell(*args, **kwargs)
File "/opt/anaconda3/lib/python3.12/site-packages/IPython/core/interactiveshell.py", line 3075, in run_cell
result = self._run_cell(
File "/opt/anaconda3/lib/python3.12/site-packages/IPython/core/interactiveshell.py", line 3130, in _run_cell
result = runner(coro)
File "/opt/anaconda3/lib/python3.12/site-packages/IPython/core/async_helpers.py", line 128, in _pseudo_sync_runner
coro.send(None)
File "/opt/anaconda3/lib/python3.12/site-packages/IPython/core/interactiveshell.py", line 3334, in run_cell_async
has_raised = await self.run_ast_nodes(code_ast.body, cell_name,
File "/opt/anaconda3/lib/python3.12/site-packages/IPython/core/interactiveshell.py", line 3517, in run_ast_nodes
if await self.run_code(code, result, async_=asy):
File "/opt/anaconda3/lib/python3.12/site-packages/IPython/core/interactiveshell.py", line 3577, in run_code
exec(code_obj, self.user_global_ns, self.user_ns)
File "/var/folders/h9/2zpp40gn24l3ddb3fnkq7fzh0000gn/T/ipykernel_50301/2443383685.py", line 18, in <module>
result = fx.optim.optimize_dataset_optical_module(
File "/Users/liam/metasurface/fouriax/src/fouriax/optim.py", line 752, in optimize_dataset_optical_module
params_result = optimize_dataset_params(
File "/Users/liam/metasurface/fouriax/src/fouriax/optim.py", line 584, in optimize_dataset_params
epoch_batches = iter_batches(epoch)
File "/Users/liam/metasurface/fouriax/src/fouriax/optim.py", line 530, in iter_batches
return batch_iter_any(train_data, epoch)
File "/Users/liam/metasurface/fouriax/src/fouriax/optim.py", line 727, in batch_iter_fn
return iter_minibatches(
File "/Users/liam/metasurface/fouriax/src/fouriax/optim.py", line 200, in iter_minibatches
from sklearn.utils import shuffle as sklearn_shuffle
File "/opt/anaconda3/lib/python3.12/site-packages/sklearn/__init__.py", line 84, in <module>
from .base import clone
File "/opt/anaconda3/lib/python3.12/site-packages/sklearn/base.py", line 19, in <module>
from .utils._estimator_html_repr import _HTMLDocumentationLinkMixin, estimator_html_repr
File "/opt/anaconda3/lib/python3.12/site-packages/sklearn/utils/__init__.py", line 11, in <module>
from ._chunking import gen_batches, gen_even_slices
File "/opt/anaconda3/lib/python3.12/site-packages/sklearn/utils/_chunking.py", line 8, in <module>
from ._param_validation import Interval, validate_params
File "/opt/anaconda3/lib/python3.12/site-packages/sklearn/utils/_param_validation.py", line 14, in <module>
from .validation import _is_arraylike_not_scalar
File "/opt/anaconda3/lib/python3.12/site-packages/sklearn/utils/validation.py", line 26, in <module>
from ..utils._array_api import _asarray_with_order, _is_numpy_namespace, get_namespace
File "/opt/anaconda3/lib/python3.12/site-packages/sklearn/utils/_array_api.py", line 11, in <module>
from .fixes import parse_version
File "/opt/anaconda3/lib/python3.12/site-packages/sklearn/utils/fixes.py", line 24, in <module>
import pandas as pd
File "/opt/anaconda3/lib/python3.12/site-packages/pandas/__init__.py", line 49, in <module>
from pandas.core.api import (
File "/opt/anaconda3/lib/python3.12/site-packages/pandas/core/api.py", line 1, in <module>
from pandas._libs import (
File "/opt/anaconda3/lib/python3.12/site-packages/pandas/_libs/__init__.py", line 17, in <module>
import pandas._libs.pandas_datetime # noqa: F401 # isort: skip # type: ignore[reportUnusedImport]
---------------------------------------------------------------------------
TypeError Traceback (most recent call last)
File ~/metasurface/fouriax/src/fouriax/optim.py:527, in optimize_dataset_params.<locals>.iter_batches(epoch)
526 try:
--> 527 return batch_iter_any(train_data, epoch, rng)
528 except TypeError:
TypeError: optimize_dataset_optical_module.<locals>.batch_iter_fn() takes 2 positional arguments but 3 were given
During handling of the above exception, another exception occurred:
ImportError Traceback (most recent call last)
File /opt/anaconda3/lib/python3.12/site-packages/numpy/core/_multiarray_umath.py:46, in __getattr__(attr_name)
41 # Also print the message (with traceback). This is because old versions
42 # of NumPy unfortunately set up the import to replace (and hide) the
43 # error. The traceback shouldn't be needed, but e.g. pytest plugins
44 # seem to swallow it and we should be failing anyway...
45 sys.stderr.write(msg + tb_msg)
---> 46 raise ImportError(msg)
48 ret = getattr(_multiarray_umath, attr_name, None)
49 if ret is None:
ImportError:
A module that was compiled using NumPy 1.x cannot be run in
NumPy 2.4.1 as it may crash. To support both 1.x and 2.x
versions of NumPy, modules must be compiled with NumPy 2.0.
Some module may need to rebuild instead e.g. with 'pybind11>=2.12'.
If you are a user of the module, the easiest solution will be to
downgrade to 'numpy<2' or try to upgrade the affected module.
We expect that some modules will need time to support NumPy 2.
batch=001/025 val_loss=0.088608 train_loss=0.087333 epoch_time=3.40s elapsed=3.40s [best]
batch=002/025 val_loss=0.087348 train_loss=0.087105 epoch_time=0.78s elapsed=4.18s [best]
batch=003/025 val_loss=0.086988 train_loss=0.083295 epoch_time=0.79s elapsed=4.97s [best]
batch=004/025 val_loss=0.086465 train_loss=0.084680 epoch_time=0.77s elapsed=5.74s [best]
batch=005/025 val_loss=0.085996 train_loss=0.085901 epoch_time=0.78s elapsed=6.52s [best]
batch=006/025 val_loss=0.085615 train_loss=0.085540 epoch_time=0.82s elapsed=7.34s [best]
batch=007/025 val_loss=0.085251 train_loss=0.083001 epoch_time=0.88s elapsed=8.21s [best]
batch=008/025 val_loss=0.084874 train_loss=0.086497 epoch_time=0.82s elapsed=9.04s [best]
batch=009/025 val_loss=0.084444 train_loss=0.083390 epoch_time=0.82s elapsed=9.86s [best]
batch=010/025 val_loss=0.083917 train_loss=0.082050 epoch_time=0.80s elapsed=10.67s [best]
batch=011/025 val_loss=0.083190 train_loss=0.082243 epoch_time=1.31s elapsed=11.98s [best]
batch=012/025 val_loss=0.082329 train_loss=0.077434 epoch_time=0.73s elapsed=12.71s [best]
batch=013/025 val_loss=0.081346 train_loss=0.081626 epoch_time=0.77s elapsed=13.48s [best]
batch=014/025 val_loss=0.080279 train_loss=0.076347 epoch_time=0.74s elapsed=14.22s [best]
batch=015/025 val_loss=0.079087 train_loss=0.075800 epoch_time=0.95s elapsed=15.17s [best]
batch=016/025 val_loss=0.077962 train_loss=0.077047 epoch_time=0.75s elapsed=15.91s [best]
batch=017/025 val_loss=0.077178 train_loss=0.074858 epoch_time=0.87s elapsed=16.79s [best]
batch=018/025 val_loss=0.076232 train_loss=0.077709 epoch_time=0.76s elapsed=17.54s [best]
batch=019/025 val_loss=0.075668 train_loss=0.074983 epoch_time=0.74s elapsed=18.28s [best]
batch=020/025 val_loss=0.075256 train_loss=0.077808 epoch_time=0.74s elapsed=19.02s [best]
batch=021/025 val_loss=0.074825 train_loss=0.071451 epoch_time=0.79s elapsed=19.81s [best]
batch=022/025 val_loss=0.074330 train_loss=0.075788 epoch_time=0.75s elapsed=20.56s [best]
batch=023/025 val_loss=0.073944 train_loss=0.074169 epoch_time=0.76s elapsed=21.32s [best]
batch=024/025 val_loss=0.073655 train_loss=0.071617 epoch_time=0.76s elapsed=22.08s [best]
batch=025/025 val_loss=0.073392 train_loss=0.073145 epoch_time=0.78s elapsed=22.86s [best]
6 Evaluation#
We test the learned filter on a separate held-out scene, then compare the resulting output with the desired edge map. We also extract the learned phase itself so it can be compared against the analytical spiral-phase solution.
final_phase = np.asarray(2.0 * jnp.pi * jax.nn.sigmoid(result.params_result.best_params))
test_scene = make_test_scene(grid)
test_target = edge_target(test_scene)
test_out = np.asarray(measure_scenes(result.best_module, test_scene, grid, spectrum)[0])
test_out_n = test_out / np.max(test_out)
cc = float(np.corrcoef(test_out_n.ravel(), np.asarray(test_target).ravel())[0, 1])
print(f"Test-scene correlation: {cc:.4f}")
spiral = np.asarray(analytical_spiral_phase(grid))
Test-scene correlation: 0.6698
7 Plot Results#
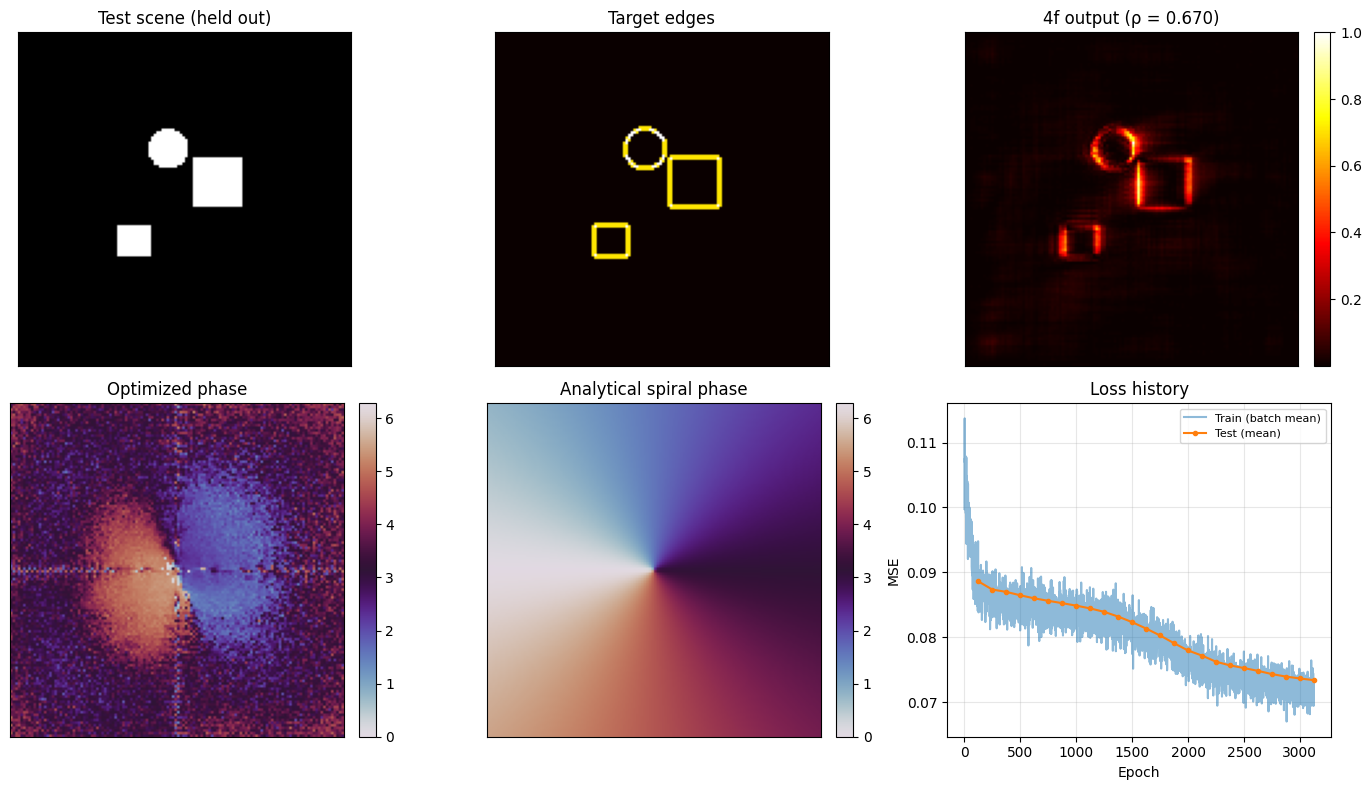
The main checks are both functional and structural:
functionally, does the held-out output look like an edge map?
structurally, does the learned phase resemble the known vortex filter?
The known optimal phase-only Fourier filter for isotropic edge detection is the vortex (spiral) phase:
where \((x, y)\) are centred coordinates in the Fourier plane (DC at the optical axis). We compare the optimised phase to this analytical solution visually.
This closes the loop on the notebook’s main claim: the 4f system can learn a classical Fourier-plane image-processing filter from data rather than having it specified by hand.
if PLOT:
fig, axes = plt.subplots(2, 3, figsize=(14, 8))
axes[0, 0].imshow(np.asarray(test_scene), cmap="gray")
axes[0, 0].set_title("Test scene (held out)")
axes[0, 1].imshow(np.asarray(test_target), cmap="hot")
axes[0, 1].set_title("Target edges")
im = axes[0, 2].imshow(test_out_n, cmap="hot")
axes[0, 2].set_title(f"4f output (ρ = {cc:.3f})")
fig.colorbar(im, ax=axes[0, 2], fraction=0.046, pad=0.04)
im_o = axes[1, 0].imshow(final_phase, cmap="twilight", vmin=0, vmax=2 * np.pi)
axes[1, 0].set_title("Optimized phase")
fig.colorbar(im_o, ax=axes[1, 0], fraction=0.046, pad=0.04)
im_s = axes[1, 1].imshow(spiral, cmap="twilight", vmin=0, vmax=2 * np.pi)
axes[1, 1].set_title("Analytical spiral phase")
fig.colorbar(im_s, ax=axes[1, 1], fraction=0.046, pad=0.04)
axes[1, 2].plot(train_history, alpha=0.5, label="Train (batch mean)")
test_steps, test_vals = zip(*test_history, strict=True)
axes[1, 2].plot(test_steps, test_vals, "o-", markersize=3, label="Test (mean)")
axes[1, 2].set_title("Loss history")
axes[1, 2].set_xlabel("Epoch")
axes[1, 2].set_ylabel("MSE")
axes[1, 2].legend(fontsize=8)
axes[1, 2].grid(alpha=0.3)
for ax in axes.flat:
if ax.images:
ax.set_xticks([])
ax.set_yticks([])
fig.tight_layout()
ARTIFACTS_DIR.mkdir(parents=True, exist_ok=True)
fig.savefig(PLOT_PATH, dpi=150)
plt.show()
print(f"saved: {PLOT_PATH}")
saved: /Users/liam/metasurface/fouriax/examples/artifacts/4f_edge_optimization.png